
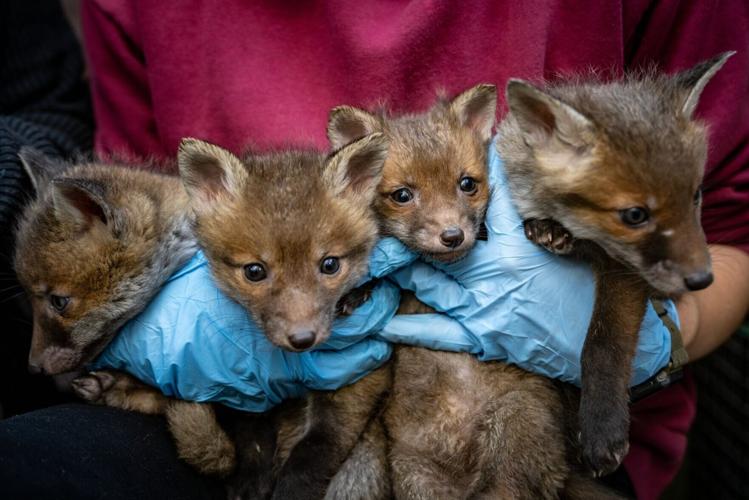
Fox cubs. (James Linsell Clark via SWNS)
Foxes, crows and magpies could provide an "early warning system" to monitor antibiotic resistance, suggests a new study.
Wildlife including the bin-dipping mammals as well as carrion birds can alert us to the spread of highly drug resistant bacteria into unexposed ecosystems, say scientists.
Antimicrobial resistance, or AMR, especially resistance against drugs critically important for human medicine, is an increasing health issue.
Scientists say one group of essential antibiotics are third-generation cephalosporins (3GCs), which are used to treat pneumonia, sepsis, meningitis and other potentially deadly infections.
Resistance to 3GCs is conferred by enzyme-encoding genes which can spread rapidly between bacteria.
Experts say one group of bacteria — sometimes labeled ESKAPE — is particularly resistant to antibiotics and can "escape" antibacterial agents.
One bacterium of that group, Klebsiella pneumoniae — which can cause severe infections in humans, has spread far beyond places and systems directly exposed to antibiotics, according to the new study.

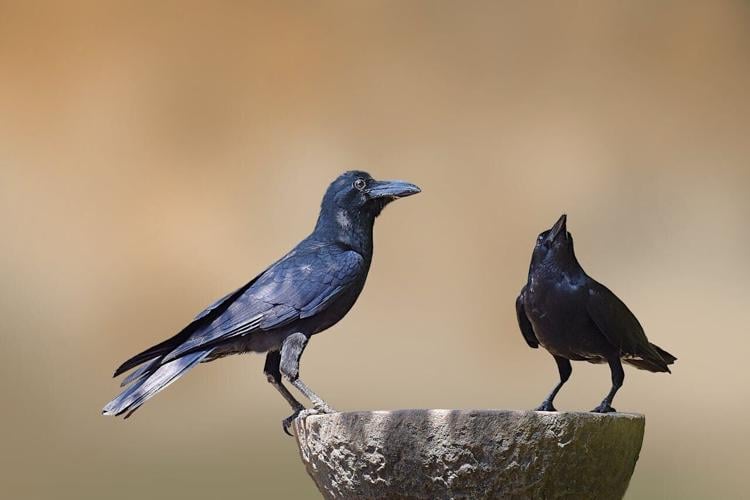
(Photo by Shilpesh Patil via Pexels)
Mauro Conter, from the University of Parma in Italy, said: "We isolated a high-risk ST307 clone of K. pneumoniae and NDM-5 carbapenemase, an enzyme variant that can inactivate antibiotics, from wildlife living far from human activity.
"This confirms the role of wildlife as reservoirs of clinically relevant resistance, which means that wildlife surveillance could provide an early warning system of resistance spreading beyond clinical settings."
The research team examined almost 500 fecal samples from wildlife.
Of the samples, 184 came from red foxes, 209 from crows and magpies, and 100 from several species of waterbirds.
The researchers used samples from the animals as they move through urban, rural, and wild areas alike, and "collect" AMR across ecosystems and regions without receiving antibiotics themselves.
Conter says foxes contribute to short-range AMR dissemination on the ground, whereas birds can spread resistance across long distances by air.
Samples were collected from animals' guts and tested for Klebsiella spp., a genus of bacteria including K. pneumoniae and other opportunistic pathogens.

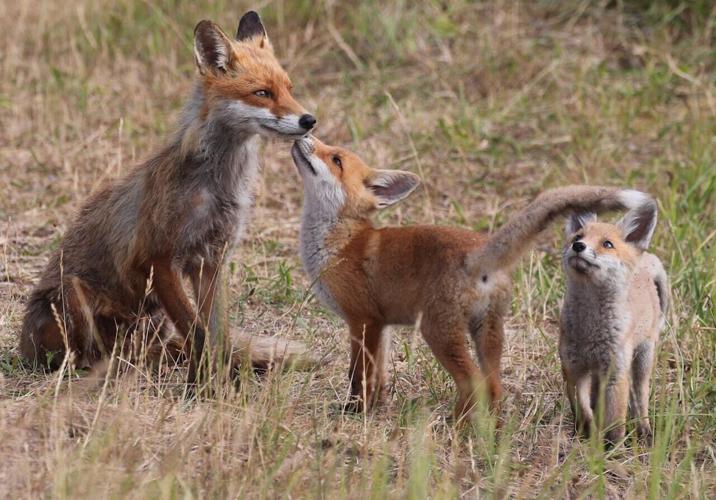
(Photo by Steffi Wacker via Pexels)
Klebsiella spp. can produce carbapenemases, which are enzymes that neutralize carbapenem antibiotics which are often "last-resort" antibiotics for treating severe infections caused by multidrug-resistant bacteria.
The research team found Klebsiella spp. bacteria in 32 samples.
Bacterial species belonging to certain genera were more or less prevalent in certain animals, with K. pneumoniae being present in waterbirds and foxes.
The bacterium was discovered in 2% of samples, according to the findings published in the journal Frontiers in Microbiology.
Conter said: "Even a 2% prevalence in wildlife represents environmental contamination by high-risk clones. K. pneumoniae readily spills over through water and waste routes, creating a continuous human-animal-environment resistance cycle."
The research team also found that K. pneumoniae isolates in the study exhibited higher levels of resistance against nearly all antibiotic classes compared to surveillance data from 2024.
The team say the trend is not surprising, but predictable across humans, animals, and the environment.
Conter said: "Our study showed that wildlife resistance exceeds clinical rates.

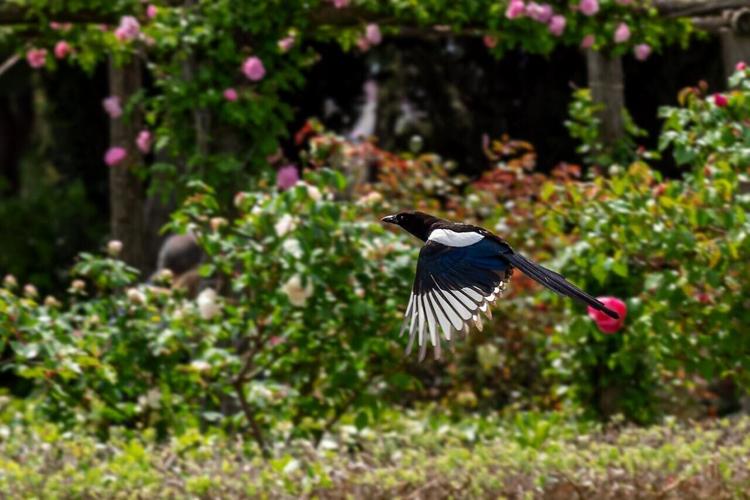
(Photo by Manuel Torres Garcia via Pexels)
"100% of K. pneumoniae isolates from wildlife in our study were resistant to 3GCs.
Compared to this, only 19.6% of K. pneumoniae isolates from human patients in Italy were resistant to 3GCs, according to the latest European Centre for Disease Prevention and Control surveillance data."
Resistance to fluoroquinolones — a different group of antibiotics used to treat severe urinary tract infections (UTIs) and pneumonia — also was at 100% in isolates from wildlife, compared to 17.4% in isolates from human patients.
To combat the spread of AMR bacteria into ecosystems that are not directly exposed to antibiotics, the researchers say we need to reduce antibiotic pollution of wastewater, improve sewage treatment, and promote more prudent use of antimicrobials in livestock.
They said that also includes restricting critically important antibiotics to human medicine.
The team pointed out that their study was not designed to identify direct transmission links between wildlife and humans, and that the prevalence of resistance rates may have been underestimated.
Due to the sampling approach, the researchers said the isolated bacteria may not represent the true diversity of bacteria in the environment.
They say larger studies that link human to animal to environment transmission — both on national and international scales — are needed, but are challenging to implement.
Conter said: "What we see is a complex problem that requires 'one health' solutions addressing antibiotic pollution, climate-driven wildlife behavioral changes, and bacterial population dynamics."
He added: "Our data justify routine wildlife AMR monitoring as a public health early warning system, guiding environmental interventions before resistance reaches clinical settings."














(0) comments
Welcome to the discussion.
Log In
Keep it Clean. Please avoid obscene, vulgar, lewd, racist or sexually-oriented language.
PLEASE TURN OFF YOUR CAPS LOCK.
Don't Threaten. Threats of harming another person will not be tolerated.
Be Truthful. Don't knowingly lie about anyone or anything.
Be Nice. No racism, sexism or any sort of -ism that is degrading to another person.
Be Proactive. Use the 'Report' link on each comment to let us know of abusive posts.
Share with Us. We'd love to hear eyewitness accounts, the history behind an article.